Figure 1
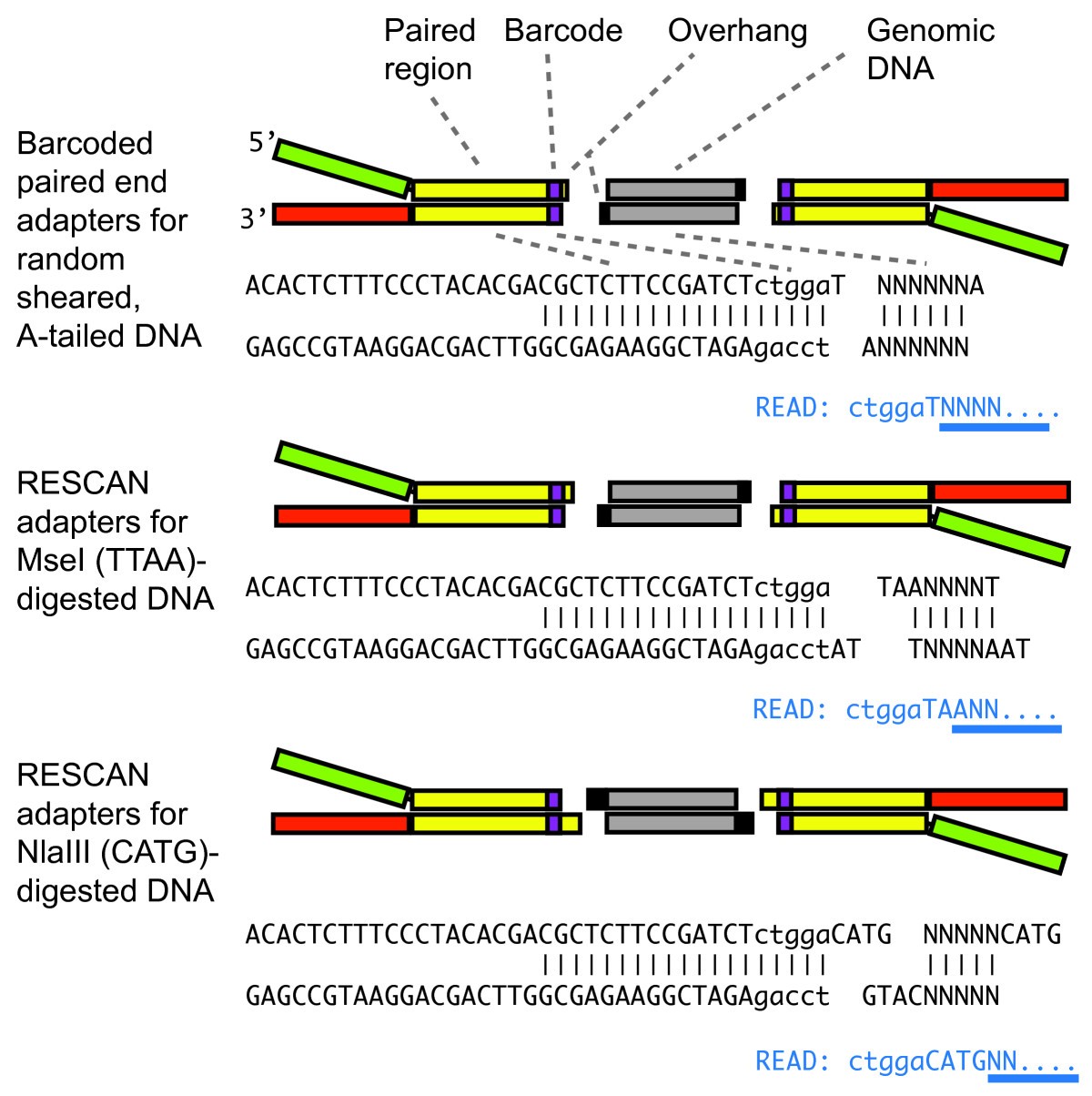
Structure of barcoded adapters used for RESCAN. The RESCAN adapters leverage the Y-adapter system used for standard Illumina sequencing libraries in which random-sheared, A-tailed insert DNA (grey boxed regions or NNN) is ligated to T-overhang formed by the paired adapters (top). The Y-adapter is formed by two oligonucleotides. A sequence barcode (lower case) is included adjacent to the end. For ligation to restriction enzyme-formed overhangs, the required extension is incorporated in the appropriate oligonucleotide of the adapter. Below each paired adapter sequence the beginning of the resulting sequence read is shown in blue, with the nucleotides that are not fixed, i.e. not part of the adapter, barcode and overhang, underlined in blue. The barcode length used in the early method-refining part of this work was of four bases. Five bases is the preferred length at the time of writing this paper because the first five cycles of Illumina HiSeq platform require random and similarly weighted base composition.