Figure 2
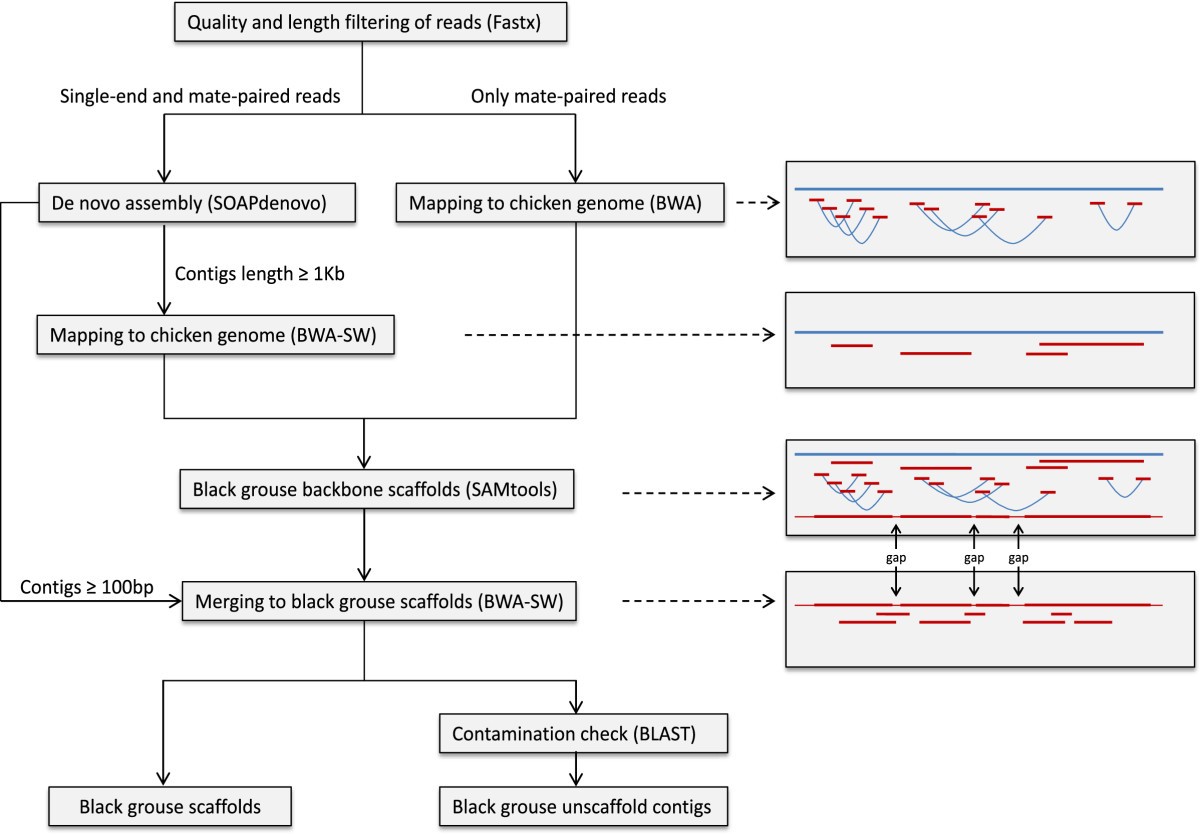
Flow chart of our reference guided genome assembly pipeline. All reads were first de-novo assembled. In the second step both long contigs and original mate-paired reads were mapped to the chicken reference genome and merged to produce the black grouse backbone scaffolds. Finally gap-filling was performed by mapping all contigs (of at least 100 bp) back to the backbone scaffolds, producing the draft genome assembly. In addition all contigs (at least 200 bp long) not mapping to the backbone scaffolds were added to the assembly after removing sequences arising from possible contamination using a BLAST approach. For more details about the procedures please see the Methods section. In the figures to the right the chicken reference genome is indicated by the blue line while reads, contigs and scaffolds from the black grouse are shown in red. The light red parts of the final scaffold indicate regions with gaps (Ns) in the black grouse sequence. Within brackets in the boxes to the left are the software used for each different stage of the assembly process.