Fig. 1
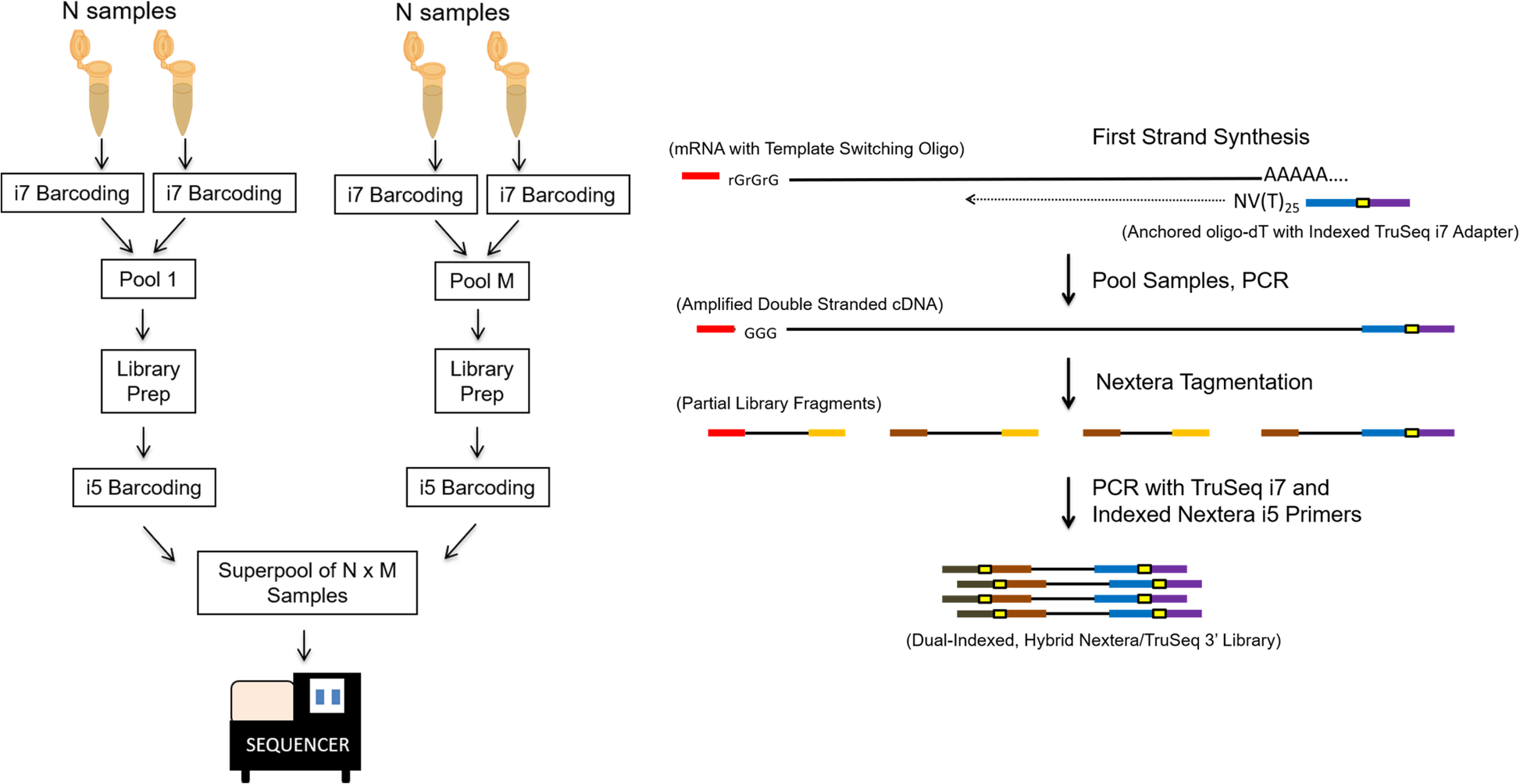
A schematic representation of the 3’Pool-seq protocol. The use of anchored oligo-dT primers with standard indexed TruSeq i7 adapter overhangs for first strand synthesis allows immediate pooling of multiple samples after reverse transcription. Within a pool, each sample can be uniquely identified by the TruSeq i7 index. Once pooled, purification, PCR, and Nextera tagmentation reagents are used to generate cDNA fragments. A second PCR step using standard TruSeq i7 and indexed Nextera i5 adapters allows selective amplification of only 3′-end cDNA fragments and barcoding of each sample pool with a standard Nextera i5 index. The final product is a dual-indexed hybrid Nextera/TruSeq 3′-library where the i5 Nextera index serves as the pool index, and the i7 TruSeq index serves as the sample index within a pool. Multiple indexed library pools can be further quantified and combined in equal proportions into a superpool for sequencing