- Research article
- Open access
- Published:
Quantitative assessment of relationship between sequence similarity and function similarity
BMC Genomics volume 8, Article number: 222 (2007)
Abstract
Background
Comparative sequence analysis is considered as the first step towards annotating new proteins in genome annotation. However, sequence comparison may lead to creation and propagation of function assignment errors. Thus, it is important to perform a thorough analysis for the quality of sequence-based function assignment using large-scale data in a systematic way.
Results
We present an analysis of the relationship between sequence similarity and function similarity for the proteins in four model organisms, i.e., Arabidopsis thaliana, Saccharomyces cerevisiae, Caenorrhabditis elegans, and Drosophila melanogaster. Using a measure of functional similarity based on the three categories of Gene Ontology (GO) classifications (biological process, molecular function, and cellular component), we quantified the correlation between functional similarity and sequence similarity measured by sequence identity or statistical significance of the alignment and compared such a correlation against randomly chosen protein pairs.
Conclusion
Various sequence-function relationships were identified from BLAST versus PSI-BLAST, sequence identity versus Expectation Value, GO indices versus semantic similarity approaches, and within genome versus between genome comparisons, for the three GO categories. Our study provides a benchmark to estimate the confidence in assignment of functions purely based on sequence similarity.
1. Background
Large-scale genome sequencing projects have discovered many new proteins. Of all the proteins whose sequences are known, functions have been experimentally determined for only a small percentage [1]. Annotation of a genome involves assignment of functions to proteins in most cases on the basis of sequence similarity. Protein function assignments based on postulated homology as recognized by sequence identity or significant expectation value of alignment are used routinely in genome analysis. Over the past years, many computational methods [2–11] have been developed to predict function through identifying sequence similarity between a protein of unknown function and one or more proteins with experimentally characterized or computationally predicted functions. However, it is widely recognized that functional annotations should be transferred with caution, as the sequence similarity does not guarantee evolutionary or functional relationship. In addition, if a protein is assigned an incorrect function in a database, the error could carry over to other proteins for which functions are inferred by sequence relationship to the protein with errant function assignment [12–14].
Despite the central role that sequence comparison programs play in functional annotation, a thorough analysis of the quality of methods based on a large-scale dataset has not been performed. Improvements in the sensitivity of sequence comparison algorithms have reached the point that proteins with previously undetectable sequence relationship, for instance with 10–15% identical residues, may be classified as similar [15]. On the other hand, alignments are more likely to be correct for higher levels of pairwise sequence identity; and are less likely to be correct in the so-called "twilight zone", where the sequence similarity is low [16]. An estimate of the expectation value of an alignment provides a good assessment for whether the two aligned proteins are homologous [17]. Nevertheless, prediction of protein function from sequence is a difficult problem, because not only sequence similarity does not guarantee homology, but also homologous proteins often have different functions [18, 19]. In particular, when two proteins are distantly related, there is no good indicator to reliably assess whether they are homologous or not. Figure 1 shows the number of unique orthologous pairs between the yeast Saccharomyces cerevisiae and Arabidopsis thaliana acquired from the Website of Clusters of Orthologous Groups of proteins (COGs) [37]. The COG pairs distribute in a broad range of sequence identity and expectation value. It is clear that neither percentage of sequence identity nor expectation value can give a complete insight into the relationship between the two proteins. Towards this we wish to study the detailed quantitative relationship in terms of functions and relate it with sequence identity and expectation value intervals.
A number of studies in sequence-function relationship have been carried out. Shah et al. [20] showed that many EC (Enzyme Commission) classes could not be perfectly discriminated by sequence similarity at any threshold. Pawlowski et al. [15] have studied the relation between sequence similarity and functional similarities based on the EC classification for the E. coli genome. However, this study is limited only to within genome comparisons and lacks any analysis based on inter-genome comparisons. Devos et al. [21] have studied the complexity in transferring function between similar sequences. Their study shows that binding site, keywords, and functional class annotations are less conserved than EC numbers, and all of them in turn are less conserved than protein structure. Wilson et al. showed that percent identity in sequence alignment is more effective at quantifying functional conservation of their simple classification of SCOP domains than modern probabilistic scores [22]. However, all these studies did not use a broad definition of functions for a systematic large-scale analysis. In this paper, we will build a comprehensive and systematic benchmark for the sequence-function relationship using four model organisms (Arabidopsis thaliana, Saccharomyces cerevisiae, Caenorrhabditis elegans, and Drosophila melanogaster) and controlled vocabularies of function annotation terms in the Gene Ontology [38] from three different perspectives, i.e., biological process, molecular function, and cellular component.
2. Results and discussion
The sequence comparisons within and across the four genomes provide a global view on the relationship between sequence similarity and function similarity. Figure 2 shows a consistent correlation between function similarity of biological process at different GO index levels and the Expectation Values (E-values) of sequence alignment using BLAST [23]. There is also a higher functional similarity for the lower GO Index levels in comparison to the higher Index levels. In particular, at levels 1 and 2, the function similarity reaches very high even when the sequence similarity is insignificant. This is mainly due to the fact that many more genes can be found under a GO Index of lower level than of higher level, and hence, there is a higher chance for two randomly picked genes to share the GO Index at the lower level (see Figure 9 and related discussion).
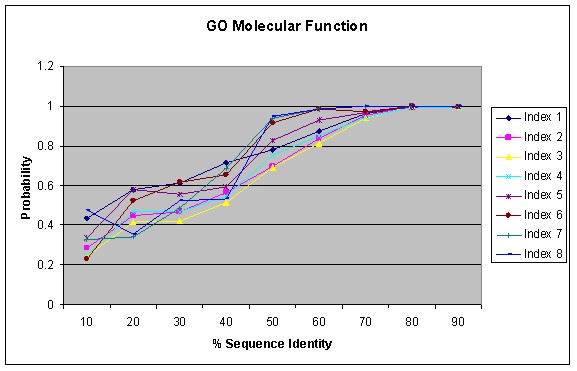
Figure 3 shows the result of function similarity with respect to sequence identity as identified by the BLAST for GO Biological Process annotations. It shows that probability of functional conservation increases with increasing sequence conservation. A similar trend is observed in different GO Index levels as in Figure 2. The probability is based on the number of pairs sharing the same function at a certain index level against the total pairs having any functions at the respective index level for a given sequence similarity interval. Such per index probability may sometimes result in higher probability for higher index levels (probably due to limited sample size) and lead to the cross-over between curves from various index levels. Interestingly, high sequence identity is a better indicator of function similarity than significant E-value as used in Figure 2. If two proteins have sequence identity more than 70%, they have about 90% probability or more to share the same biological process for GO index levels 1–8. On the other hand, E-value depends on many factors, in particular the lengths of the two proteins. For large proteins with homologous relationship, the E-value tends to be more significant for computational identification of the homology relationship, but their sequence identity can be very weak and their functional relationship may be remote. Figures 4 and 5 show similar results as Figure 3 for GO Molecular Function and GO Cellular Component Annotations, respectively. The result is similar to that observed by Pawlowski et al. in their studies on enzymes based on the E. coli genome [15] and by Wilson et al [22] who use FLY+ENZYME classification SCOP domains, MIPS and GenProtEC to study sequence and functional conservation.
Functional conservation measures from GO annotations based on computational techniques such as electronic annotation based on sequence similarity has a behavioral pattern completely different from Figures 3, 4 and 5. When GO Biological process annotations are made based on evidences from experimental validations (Figure 6A), such proteins tend to conserve and share functions with higher probability for pairs with high sequence identity as compared to pairs with remote sequence similarity. In all cases, when a pair of proteins share sequence identity 30% or less, the chance for them to share any of the three GO categories at high levels is about 50% or less. However, this pattern is lost when annotations are made purely based on computational techniques (Figure 6B) and the functions are conserved with almost equal probabilities irrespective of the sequence conservation. This depicts the difference in the quality of these two annotations, and indicates that many annotations based on computational techniques may be incorrect. Some of these incorrect annotations could be due to over-extension of functional details when inferring a query protein from a protein hit with known function. For example, a protein in two-component signal transduction system (GO:0000160) could be predicted to cell surface receptor linked signal transduction (GO:0007166), although both proteins are in signal transduction (GO:0007165). The trend stands true for both Molecular Function and Cellular Component (data not shown).
We also compared the SubLoc predicted localizations for all the proteins across genomes. Figure 7 shows the localization similarity versus the sequence similarity in terms of E-value and percentage of sequence identity for intra-genome comparisons within four genomes. In this case the localization is measured by five types as described in Section 4.4, instead of the GO Cellular Component Annotation, a detailed level that no existing software can predict reliably. Subcellular localization conservation shows similar results when compared in terms of E-value or sequence identity. Inter-genome comparisons based on the predicted subcellular localizations also behave in a manner similar to the intra-genome comparisons (data not shown). It is interesting to note that the behavior of the curves of the four genomes is similar in respect of E-value (Figure 7A). On the other hand, the behavior of the curves of the four genomes shows the difference in respect to the sequence identity (Figure 7B), in particular, Caenorrhabditis elegans shows significantly more divergence in localization under the same sequence identity than the other three genomes.
A. Relation between E-value intervals (negative logarithmic with base 10) of seq uence similarity and similarity in SubLoc predicted localization of proteins within the same genomes using FASTA. B. Relation between percentage of sequence similarity and similarity in SubLoc predicted localization of proteins within the same genomes using BLAST.
We also calculated functional similarity in terms of semantic similarity between the functional annotation terms of GO (see section 4.3) using BLAST. Figure 8 shows the relationship between semantic similarity and sequence identity for all four genomes combined for all three Ontologies. The semantic similarity measures remove the bias seen between different levels of indices of the ontology. Figure 9 shows the relationship between remote homologs using PSI-BLAST [24] and semantic similarity. For many of the PSI-BLAST pairs, the sequence identity is below 30%. Hence, we focus the sequence-function relationship based on E-value, instead of sequence identity.
We have also computed results as described above for any random pairs with known function annotation. Then, we calculated a normalized ratio of function similarity in terms of sequence identity by comparing the results in Figures 3 through 5 against similar results from random pairs. Figure 10 shows the normalized ratio results for GO Biological Process, Molecular Function and Cellular Component Annotations in subplots A, B and C, respectively. Our results clearly show that the normalized ratio increases for higher sequence identity intervals as well as higher levels of shared GO Indices, highlighting the higher chance of functional conservation over randomly chosen pairs for these groups. GO annotations for Index level 3 and above are very informative as the probability of correct functional assignment based on sequence similarity is significantly above random. Figure 10D shows normalized results for all three annotations using PSI-BLAST in subplot D. It indicates that PSI-BLAST has substantial enrichment of function assignment for function prediction. This may be because PSI-BLAST utilizes multiple sequence profiles that enhance the recognition of the sequence-function relationship.
Relation between percentage of sequence similarity and functional similarity for GO (A) Biological Process, (B) Molecular Function and (C) Cellular Component Annotations within the same genomes using BLAST and (D) for all annotations using PSI-BLAST respectively, in the form of normalized ratio of pms(t1, t2), which is the probability of the minimum subsumer for terms t1 and t2 (see section 4.3).
Figure 11 and 12 show results similar to Figure 10 for normalized ratio against random pairs for inter-genome sequence similarity comparisons between each yeast protein and the other three genomes, for GO Biological Process and Molecular Function Annotations, respectively. It appears that the trend of biological process in inter-genome comparison is similar to the one in intra-genome comparison, while the trend of molecular function in inter-genome comparison is very diverse. This suggests that many homologous genes may have evolved into different molecular functions in different genomes.
Relation between percentage of sequence similarity and functional similarity for the GO Molecular Function Annotations for inter-genome comparison of yeast ORFs against others using BLAST, in the form of normalized ratio. Data points with a sample size less than 10 gene pairs are not sure, as the statistics is not significant.
3. Conclusion
It has been long recognized that genome annotations using computational methods produce many false function assignments. Many of these methods have been applied to function prediction. They often provide valuable hypotheses, but none are perfect. As a result, it is known that many databases contain incorrect function assignments, and these erroneous assignments propagate from one database to another. Nevertheless, up until now there has been no systematic study for this critical issue. The question whether two proteins are functionally similar is very complex to answer. Function is a very complex notion involving many different aspects including chemical, biochemical, cellular, organism mediated, and developmental processes. Qualitatively it is expected that with higher sequence similarity, the two proteins are more likely to have related functions. However, quantitatively the relationship between function similarity at the different categories and sequence similarity has not been studied deeply. Such a quantitative study is fundamentally important, as it can provide assessment of gene function prediction quality and insights into the underlying mechanisms of new evolving functions through changes in sequence [25, 26].
Our study confirms that sequence comparison often provides good suggestions for gene functions or related functions. These suggestions serve as useful hypotheses for further experimental work to confirm, refine or refute the predictions. Such a process can substantially increase the speed of biological knowledge discovery. On the other hand, when assigning function based purely on similarity to proteins of known function (as annotated in databases), it is important to be aware of incomplete or wrong annotations. Given the value of computational function annotation, our study also shows that a significant portion of gene annotations of biological process, molecular function, and cellular component based solely on sequence similarity, in particular, when the sequence similarity is low, are unreliable. Our study also provides a numerical benchmark for the extent to which one can trust computational annotation. It is possible that a confidence score can be derived from our study for any annotation based on sequence similarity. With this score in the annotation file, the user can have a better insight about the quality of the annotations. Furthermore, our analyses highlights the different sequence-function relationships identified from BLAST versus PSI-BLAST, sequence identity versus Expectation value, GO indices versus semantic similarity approaches and within genome versus between genome comparisons, for the three GO classification types.
There are some limitations in our current study. Our study can only reflect certain aspect of protein function. Protein function variations may result from factors other than sequence, such as alternative splicing and post-translational modification, and our method does not address these factors. Another limitation is that when we assess gene function prediction, we only consider one hit at a time in a database. In many cases, sequence comparison yields multiple hits for one query protein and these hits may have different functions. In our future study, we will develop a new method to assess the function prediction for a query protein by combining the functions of multiple hits while considering the dependence among these functions and the E-values of the hits.
4. Methods
4.1 Protein sequence databases
We selected the genomes of Arabidopsis thaliana, Saccharomyces cerevisiae, Caenorrhabditis elegans and Drosophila melanogaster for the study. All four genomes are well-studied model organisms in eukaryotes. The complete set of Arabidopsis thaliana protein sequences for 27,288 ORFs was acquired from The Arabidopsis Information Resource (TAIR) [39]. We also obtained proteins sequences for 21,588 Caenorrhabditis elegans ORFs, 6350 Saccharomyces cerevisiae ORFs and 13,665 Drosophila melanogaster ORFs from NCBI [40]. Table 1 lists the number of ORFs for all the four genomes whose functions are annotated based on experimental evidences or sequence similarity measures for all the three functional categories.
4.2 Protein functional classification
The Gene Ontology (GO) functional classification [27] has three functional categories, i.e., biological process, molecular function and cellular component. It is not a hierarchical tree but the directed acyclic nature of the graph can be well captured in a series of numerical numbers. We have generated a numerical GO INDEX for all three classifications individually, which represents the structure of every ontology. The deepest level of index is 13. A GO Index, as denoted by numbers, e.g. 1-4-2-29, characterizes the function of every protein. The first number corresponds to the type of functional category, e.g. 1 represents biological process, 2 represents molecular function and 3 represents cellular component. The subsequent numbers correspond to subcategories describing the type of function or localization in increasing detail. The higher the GO Index level, the more specific is the functional category the protein belongs to. Table 2 shows an example of GO indices.
We assume that the functional relationship between two proteins is reflected by the number of index levels that they share. We have demonstrated the usefulness of such an assumption in our early studies for gene function prediction [28, 29]. We acquired the GO annotations for all the genes in the four genomes and for the three functional categories from GO Website [38]. A gene can (and usually does) belong to multiple indices at various levels in the graph, as proteins may be involved in multiple functions in a cell. Different indices could correspond to the same GO term as well.
Gene Ontology annotation is based on various evidences to annotate functional categories. Towards quality control, all the plots (except for Figure 6B) presented in this paper are based on the annotations with actual experimental evidences such as IDA (inferred from direct assay), IEP (inferred from expression pattern), IGI (inferred from genetic interaction), IMP (inferred from mutant phenotype), IPI (inferred from physical interaction), RCA (inferred from reviewed computational analysis) and TAS (traceable author statement). We performed some comparisons using annotations assigned purely based on computational methods such as ISS (inferred from sequence similarity) and IEA (inferred from electronic annotation), but the plots are not presented here. We have removed the functional annotations that were purely based on evidences such as ND (no biological data available) and NAS (non-traceable author statement.
4.3 Protein functional similarity
Within each family of proteins with similar sequences, functional similarity between proteins is expressed as the number of common roots shared by their functional classification other than the first level, which represents a classification of biological process, molecular function and cellular component. In the case of proteins with multiple functional assignments, the maximum indices of overlap are considered. For example, consider a gene pair ORF1 and ORF2, both annotated proteins. Assume ORF1 has a function represented by GO INDEX 1-1-3-3-4 and ORF2 has a function 1-1-3-2. When compared with each other for the level of matching GO INDEX, they match through INDEX level 1 (1-1) and level 2 (1-1-3) and will have functional similarity equal to 2. The functional similarity defined this way can assume values from 1 to 12.
We also calculate functional similarity in terms of semantic similarity between the GO functional annotation terms [30, 31]. An example of calculating the probabilities is shown in Figure 13. To calculate semantic similarity between the protein pairs, the probability of each term assigned to the gene product is first derived. For each gene in the organism, the probability is calculated by counting the number of the descendants of an assigned GO term plus 1 (the GO term itself), divided by the total number of GO term annotations in the organism. The probability of each node increases as we go towards the root of the GO ontology, which is defined as "Biological Process" (GO:0008150), "Molecular Function" (GO:0003674) or "Cellular Component" (GO:0005575) in the three Ontologies and has a probability of 1. The semantic similarity between ontology terms is defined as:
SS (t1,t2) = -ln p ms (t1,t2)
where, p ms (t1, t2) is the probability of the minimum subsumer for terms t1 and t2. The minimum subsumer for terms t1 and t2 is defined as the common parent of the deepest GO Index level shared by t1 and t2.
4.4 Protein subcellular localization
The subcellular distribution of proteins within a proteome is useful and important to a global understanding of the molecular mechanisms of a cell. Protein localization can be seen as an indicator of its function. Localization data can be used as a means of evaluating protein information inferred from other resources. Furthermore, the subcellular localization of a protein often reveals its activity mechanism. The subcellular localization information was predicted using SubLoc [32, 33, 41]. The five main subcellular localization categories as predicted by SubLoc are Cytoplasmic, Nuclear, Mitochondrial, Transmembrane, and Extracellular. The total numbers of proteins with predicted subcellular localization are 6323 in Saccharomyces cerevisiae, 27,288 in Arabidopsis thaliana, 21,588 in Caenorrhabditis elegans, and 18,498 in Drosophila melanogaster. It is worth mentioning that the subcellular localization predictions were not based on sequence similarity.
4.5 Protein sequence similarity search
The sequence similarity search was done using tools such as BLAST [23], FASTA [34, 35] and PSI-BLAST [24]. BLAST is the most widely used sequence comparison tool, particularly for genome annotation. FASTA is more sensitive in accuracy but slower than BLAST. Both FASTA and BLAST were developed for pairwise local alignment, with heuristics used. PSI-BLAST is used to identify remote homology based on iterative BLAST searches.
We compared the sequences for within as well as between genome sequence similarities. Each protein sequence was compared against the complete set of proteins for the same genome for within genome comparisons. For between genome comparisons, a pair of similar protein pair was identified using the reciprocal search method [36], i.e., the two proteins in the pair are the best hits in each other's genome from sequence search. Intra-genome sequence comparison would reflect the sequence similarity between the paralogs; while the inter-genome comparison would partially highlight the orthologous sequence similarities.
To assess the significance of a sequence comparison, an expectation value or E-value can be calculated. This value represents the number of different alignments with the observed alignment score or better that are expected to occur in the database search simply by chance. The E-value is a widely accepted measure for assessing potential biological relationship, as it is an indicator of the probability for finding the match by chance. Smaller E-values represent more likelihood of having an underlying biological relationship. In this study, we will use both E-value and sequence identity as parameters to quantify sequence similarity. On the other hand, E-values depend on a number of computational factors, such as the length of the query protein and the size of search database. The issues prevent the E-value from being a reliable indicator for homology, as addressed in Fig. 1 and related discussions.
4.6 Availability
The data and results are publicly available at our website [42].
References
Andrade MA, Sander C: Bioinformatics: from genome data to biological knowledge. Current Opinion in Biotechnology. 1997, 8: 675-683. 10.1016/S0958-1669(97)80118-8.
Koonin EV, Bork P, Sander C: Yeast chromosome III: new gene functions. The EMBO Journal. 1994, 13: 493-503.
Casari G, Sander C, Valencia A: A method to predict functional residues in proteins. Nature Structural Biology. 1995, 2: 171-178. 10.1038/nsb0295-171.
Ouzounis C, Casari G, Sander C, Tamames J, Valencia A: Comparisons of Model Genomes. Trends in Biotechnology. 1996, 14 (B): 280-285. 10.1016/0167-7799(96)10043-3.
Schneider R, Casari G, Antoine DD, Bremer P, Schlenkrich M: GeneCrunch: Experiences on the SGI POWER CHALLENGE array with bioinformatics applications. Supercomputer 1996: Anwendungen, Architekturen, Trends. 1997, , 109-119.
Bork P, Ouzounis C, Sander C: From genome sequences to protein function. Curr Opin Struct Biol. 1994, 4: 39-403. 10.1016/S0959-440X(94)90109-0.
Bork P, Koonin EV: Predicting functions from protein sequences-where are the bottlenecks?. Nat Genet. 1998, 18: 313-318. 10.1038/ng0498-313.
Bork P, Dandekar T, Diaz-Lazcoz Y, Eisenhaber F, Huynen M, Yuan Y: Predicting function: from genes to genomes and back. J Mol Biol. 1998, 283: 707-725. 10.1006/jmbi.1998.2144.
Tamames J, Ouzounis C, Casari G, Sander C, Valencia A: EUCLID: Automatic Classification of Proteins in Functional Classes by Their Database Annotations. Bioinformatics. 1998, 14: 542-543. 10.1093/bioinformatics/14.6.542.
Andrade MA, Brown NP: "Automated genome sequence analysis and annotation". Bioinformatics. 1999, 15: 391-412. 10.1093/bioinformatics/15.5.391.
Koonin EV: "Computational genomics". Curr Biol. 2001, 11: R155-158. 10.1016/S0960-9822(01)00081-1.
Brenner SE: Errors in genome annotation. Trends Genet. 1999, 15: 132-133. 10.1016/S0168-9525(99)01706-0.
Karp PD: A protocol for maintaining multidatabase referential integrity. Pac Symp Biocomput. 1996, 438-445.
Karp P: What we do not know about sequence analysis and sequence databases. Bioinformatics. 1998, 14: 753-754. 10.1093/bioinformatics/14.9.753.
Pawlowski K, Jaroszewski L, Rychlewski L, Godzik A: Sensitive sequence comparison as protein function predictor. Pac Symp Biocomput. 2000, 42-53.
Rost B, Valencia A: Pitfalls of protein sequence analysis. Curr Opin Biotechnol. 1996, 7: 457-461. 10.1016/S0958-1669(96)80124-8.
Levitt M, Gerstein M: A unified statistical framework for sequence comparison and structure comparison. Proc Natl Acad Sci USA. 1998, 95: 5913-5920. 10.1073/pnas.95.11.5913.
Whisstock JC, Lesk AM: Prediction of protein function from protein sequence and structure. Q Rev Biophys. 2003, 36: 307-40. 10.1017/S0033583503003901.
Ponting C: Issues in predicting protein function from sequence. Brief Bioinform. 2001, 2: 19-29. 10.1093/bib/2.1.19.
Shah I, Hunter L: Predicting enzyme function from sequence: a systematic appraisal. Proc Int Conf Intell Syst Mol Biol. 1997, 5: 276-283.
Devos D, Valencia A: Practical limits of function prediction. Proteins. 2000, 41 (1): 98-107. 10.1002/1097-0134(20001001)41:1<98::AID-PROT120>3.0.CO;2-S.
Wilson CA, Kreychman J, Gerstein M: Assessing annotation transfer for genomics: quantifying the relations between protein sequence, structure and function through traditional and probabilistic scores. J Mol Biol. 2000, 297 (1): 233-49. 10.1006/jmbi.2000.3550.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ: Basic local alignment search tool. J Mol Biol. 1990, 215: 403-410.
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ: Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997, 25 (17): 3389-3402. 10.1093/nar/25.17.3389.
Thornton JM, Orengo CA, Todd AE, Pearl FM: Protein folds, functions and evolution. J Mol Biol. 1999, 293: 333-342. 10.1006/jmbi.1999.3054.
Thornton JM: From genome to function. Science. 2001, 292: 2095-2097. 10.1126/science.292.5524.2095.
The Gene Ontology Consortium. Nature Genetics. 2000, 25: 25-29. 10.1038/75556.
Joshi T, Chen Y, Becker JM, Alexandrov N, Xu D: Genome-Scale Gene Function Prediction Using Multiple Sources of High-Throughput Data in Yeast. Saccharomyces cerevisiae. 2004, OMICS: A Journal of Integrative Biology, 8 (4): 322-333.
Chen Y, Xu D: lobal Protein Function Annotation through Mining Genome-Scale Data in Yeast Saccharomyces cerevisiae. G. Nucleic Acid Research. 2004, 32: 6414-6424. 10.1093/nar/gkh978.
Lord PW, Stevens RD, Brass A, Goble CA: Investigating semantic similarity measures across the Gene Ontology: the relationship between sequence and annotation. Bioinformatics. 2003, 19 (10): 1275-83. 10.1093/bioinformatics/btg153.
Resnik P: Semantic similarity in a taxonomy: an information based measure and its application to problems of ambiguity in natural language. J Artif Intelligence Res. 1999, 11: 95-130.
Guo T, Hua S, Ji X, Sun Z: DBSubLoc: database of protein subcellular localization. Nucleic Acids Research. 2004, D122-D124. 10.1093/nar/gkh109. 32 Database
Hua S, Sun Z: Support vector machine approach for protein subcellular localization prediction. Bioinformatics. 2001, 17: 721-728. 10.1093/bioinformatics/17.8.721.
Pearson WR, Lipman DJ: Improved Tools for Biological Sequence Analysis. Natl Acad Sci USA. 1988, 85: 2444-2448. 10.1073/pnas.85.8.2444.
Pearson WR: Rapid and Sensitive Sequence Comparison with FASTP and FASTA. Methods in Enzymology. 1990, 183: 63-98.
Tatusov RL, Galperin MY, Natale DA, Koonin EV: The COG database: a tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res. 2000, 28: 33-36. 10.1093/nar/28.1.33.
Clusters of Orthologous Groups (COGs). [http://www.ncbi.nlm.nih.gov/COG/]
The Gene Ontology. [http://www.geneontology.org/]
The Arabidopsis Information Resource (TAIR). [ftp://ftp.arabidopsis.org/]
National Center for Biotechnology Information(NCBI). [ftp://ftp.ncbi.nih.gov/]
Subcellular Localization Prediction of Eukaryotic Proteins (SubLoc). [http://www.bioinfo.tsinghua.edu.cn/SubLoc/eu_predict.htm]
Data and Results website. [http://digbio.missouri.edu/sfsimilarity]
Acknowledgements
This research is supported by USDA/CSREES-2004-25604-14708 and NSF/ITR-IIS-0407204. We like to thank the anonymous reviewers for their helpful suggestions.
Author information
Authors and Affiliations
Corresponding author
Additional information
Authors' contributions
TJ contributed in the data collection, sequence alignments and generation and analysis of the results. Both TJ and DX contributed in the formulation, design and writing of the study. Both authors read and approved the final manuscript.
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under license to BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Joshi, T., Xu, D. Quantitative assessment of relationship between sequence similarity and function similarity. BMC Genomics 8, 222 (2007). https://doi.org/10.1186/1471-2164-8-222
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/1471-2164-8-222