- Research article
- Open access
- Published:
Choice of library size normalization and statistical methods for differential gene expression analysis in balanced two-group comparisons for RNA-seq studies
BMC Genomics volume 21, Article number: 75 (2020)
Abstract
Background
High-throughput RNA sequencing (RNA-seq) has evolved as an important analytical tool in molecular biology. Although the utility and importance of this technique have grown, uncertainties regarding the proper analysis of RNA-seq data remain. Of primary concern, there is no consensus regarding which normalization and statistical methods are the most appropriate for analyzing this data. The lack of standardized analytical methods leads to uncertainties in data interpretation and study reproducibility, especially with studies reporting high false discovery rates. In this study, we compared a recently developed normalization method, UQ-pgQ2, with three of the most frequently used alternatives including RLE (relative log estimate), TMM (Trimmed-mean M values) and UQ (upper quartile normalization) in the analysis of RNA-seq data. We evaluated the performance of these methods for gene-level differential expression analysis by considering the factors, including: 1) normalization combined with the choice of a Wald test from DESeq2 and an exact test/QL (Quasi-likelihood) F-Test from edgeR; 2) sample sizes in two balanced two-group comparisons; and 3) sequencing read depths.
Results
Using the MAQC RNA-seq datasets with small sample replicates, we found that UQ-pgQ2 normalization combined with an exact test can achieve better performance in term of power and specificity in differential gene expression analysis. However, using an intra-group analysis of false positives from real and simulated data, we found that a Wald test performs better than an exact test when the number of sample replicates is large and that a QL F-test performs the best given sample sizes of 5, 10 and 15 for any normalization. The RLE, TMM and UQ methods performed similarly given a desired sample size.
Conclusion
We found the UQ-pgQ2 method combined with an exact test/QL F-test is the best choice in order to control false positives when the sample size is small. When the sample size is large, UQ-pgQ2 with a QL F-test is a better choice for the type I error control in an intra-group analysis. We observed read depths have a minimal impact for differential gene expression analysis based on the simulated data.
Background
High-through RNA sequencing (RNA-seq) has been increasingly used in the studies of genomics and transcriptomics over the last decade [1, 2]. Unlike cDNA microarray technology, RNA-seq has wide applications for the identification of novel genes or transcripts, mutations, gene editing and differential gene expression [1, 3,4,5,6,7]. Recent clinical studies demonstrated the utility of RNA-seq in identifying complex disease signatures via transcriptome analysis [8, 9]. Despite this utility and importance, optimal methods for analyzing RNA-seq data remain uncertain.
For each sample in an RNA-seq experiment, millions of reads with a desired read length are mapped to a reference genome by alignment tools such as Bowtie2/TopHat2, STAR and HISAT2 [10,11,12,13,14]. The mapped reads for each gene or transcript are subsequently used to quantify its expression abundance. However, the sample read depth typically varies from one sample to another and a direct comparison of gene expression between samples cannot be performed. Thus, normalization and proper test statistics are critical steps in the analysis of RNA-seq data [15].
Normalization of RNA-seq read counts is an essential procedure that corrects for non-biological variation of samples due to library preparation, sequencing read depth, gene length, mapping bias and other technical issues [16,17,18,19,20]. This ensures proper modeling of biological variations to directly compare and accurately detect expression changes between sample groups. Currently, a number of normalization methods are available to correct for technical variations and biases. These include methods to correct for read depth and transcript lengths, most commonly formulated as RPKM (Reads Per Kilobase per Million mapped reads) and FPKM (Fragments Per Kilobase per Million mapped fragments), which have been implemented in DEGSeq and Cufflinks-CuffDiff [7, 19, 21, 22]. Other global scaling quantile normalization methods consider either a TC (per-sample total counts) [23], UQ (per-sample 75% upper quartile Q3) [17], Med (per-sample Median Q2) [23], or Q (full quantile) implemented in Aroma.light [24]. More complex methods based on a size factor imputed include RLE normalization as implemented in DESeq2/DESeq and TMM implemented in edgeR for correcting read depth bias [16, 25, 26]. Still other methods normalize by the expression of control genes such as RUV for removing unwanted technical variation across samples [17, 27], GC-content [28], or log2 transformed read counts implemented in voom-limma [24, 29]. In addition to these traditional normalization methods, two abundance estimation normalization methods have been recently developed. One is called RNA-seq by Expectation-Maximization using a directed graph model (RSEM) [30] and the other is Sailfish which is an alignment-free abundance estimation using k-mers to index and count RNA-seq reads [31]. More recently we developed a method called UQ-pgQ2 (per-gene Q2 normalization following per-sample upper-quartile global scaling at 75 percentile) for correcting library depths and scaling the reads of each gene into the similar levels across conditions [18, 32].
A number of studies have compared these normalization methods and their impact on the downstream analysis for identification of differentially expressed genes (DEGs) (Table 1). Briefly, the earliest comparison studies reported that UQ normalization followed by an exact test/LRT significantly reduced the length bias of DE from RPKM relative to quantitative Real-Time polymerase chain reaction (qRT-PCR) [17] and baySeq with UQ normalization had the highest true positive rates with low false positive rates (FPRs). The observed false discovery rate (FDR) from edgeR, DESeq and TSPM methods was higher than the expected rate of 0.05, while TSPM performs the worst when sample sizes were as small as two [33]. In contrast, Rapaport et al. reported that no single method was favorable in all comparisons. They observed that baySeq with UQ normalization was the least correlated with qRT-PCR, Cufflinks-CuffDiff had an inflated number of false positive predictions and voom-limma package had comparable performance as DESeq and edgeR [34]. Moreover, a recent study based on a Spearman correlation analysis between read counts and qRT-PCR for the two abundance estimation methods (Sailfish and RSEM) revealed that raw counts (RC) or RPKM seemed to be adequate due to inconsistent results from Sailfish and RSEM, suggesting that normalization methods are not necessary for all sequence data [35]. An extensive evaluation performed by Dillest et al. found that an exact test combined with DESeq/TMM normalization was the best for controlling the FDR below 0.05 for high-count genes while RPKM, TC and Q normalization were suggested to be abandoned [23]. Moreover, several studies summarized that DESeq was often too conservative, edgeR, NBPSeq, and EBSeq were too liberal, and voom/vst-limma had good type I error control with low power for small sample sizes [36,37,38,39]. These studies concur that DESeq is preferred for controlling the number of false positives while edgeR with TMM is slightly preferable for controlling false negatives by achieving higher sensitivity.
Since DESeq with an exact test was overly conservative, DESeq2 with a Wald test was developed for improving the sensitivity/power [25]. Subsequently, a comparison of RLE normalization from DESeq2 with other existing methods was performed by several studies (Table 1). In one of these studies, a three-group comparison calculated the area under a Receiver Operating Characteristic (ROC) curve and recommended edgeR for count data with replicates while DESeq2 with RLE normalization was recommended for data without replicates [40]. Another study reported that voom and edgeR were generally superior to other methods for controlling the FDR with replicates of 3 and 5, but voom significantly underperformed in transcript-level simulation [41]. In contrast, another study reported that TMM, RLE and MRN gave the same results for a two-condition comparison without replicates [42] while limma-Voom, NOIseq and DESeq2 had more consistent results for DEG identification [43]. A recent study using RNA-seq time course data found DESeq2 and edgeR with a pairwise comparison outperformed TC tools for short time course (< 8 time points) due to high FPRs, but they performed worse on longer time series than splineTC and maSigPro tools [44].
Taken together, these studies showed that TMM and/or RLE associated with edgeR and DESeq2 outperformed the others in terms of overall performance on sensitivity and specificity [17, 23, 33, 34, 36, 37, 39,40,41, 43, 44]. However, these studies also reported that TMM and UQ normalizations were too liberal or oversensitive, resulting in a large number of false positives, while RLE implemented in DESeq with an exact test was too conservative [23, 36, 37]. A recent study concluded that RLE/DESeq2 with a Wald test improves sensitivity compared with a previous version of RLE/DESeq with an exact test. But this comes with a trade-off for a relatively higher FPR [25]. Later studies reported that the actual FDR produced from TMM/edgeR with an exact test, and RLE/DESeq2 with a Wald test, was not controlled well in many cases [18, 23, 33, 36, 37]. Most recently, edgeR offered a quasi-likelihood (QL) F-test for testing DE genes using negative binomial generalized models, which was considered to be a preferred choice for the uncertainty in estimating the dispersion for each gene when sample sizes were small [45]. In our recent study, we found that UQ-pgQ2 normalization combined with an exact test from edegR performed slightly better than TMM and RLE in terms of FDR when using MAQC data and simulated data. However, all the methods had an inflated FDR using MAQC datasets [18]. Thus, it remains unclear which combination of normalization and test statistics can minimize the number of false positives while taking into consideration of sample size and read depth variations. While studies comparing different normalization methods have been widely reported and discussed, this issue for the evaluation of newly developed normalization and testing statistical methods has not been adequately addressed.
In this study, we evaluated the performance of two commonly used packages (DESeq2 and edgeR) with three statistical tests (exact test, QL F-test and Wald test), the three most frequently used normalization methods (RLE, TMM and UQ) and the more recently proposed two-step normalization (UQ-pgQ2). Two benchmark MAQC (Microarray Quality Control Project) datasets [34, 46, 47], five real RNA-seq datasets from The Cancer Genome Atlas (TCGA) website [48], and simulated data with varying read depths are used in this study.
Results
Statistical analysis of MAQC2 and MAQC3 for the combined methods
In our previous study, we evaluated the effect of normalization methods including DESeq, TMM, UQ-pgQ2 and UQ based on DEG analysis using two MAQC datasets and an exact test/edgeR. In this study, the effects of the Wald test/DESeq2, exact test/QL F-test from edgeR and t-test/voom-limma were used to evaluate the normalization and test statistical methods.
The number of true positive (TP) and false positive (FP) genes calculated were based on the number of DEGs identified from MAQC RNA-seq data given a nominal FDR cutoff 0.05, and the total number of TPs and true negatives (TNs) were based on qRT-PCR data. We also calculated the positive predictive value (PPV), the actual FDR, sensitivity and specificity for both datasets (Table 2). Using MAQC2 data, the analysis results show that UQ-pgQ2 combined with an exact test/edgeR has the highest specificity (85.1%) with the lowest actual FDR (0.055) while the others ranged from 37.8 to 45.3% with a FDR greater than 0.1 and slightly lower sensitivity (96.7%). An exact test/TMM has the highest sensitivity (98.5%) while the others ranged from 96.7 to 97.4%. The UQ approach performed the worst in both sensitivity and specificity, consistent with other findings [18].
While using a Wald test, the results show that UQ-pgQ2 outperformed the others with the highest specificity (66.9% compared to the others from 43.9 to 46.0%) and a slightly higher sensitivity (98.7% compared to the others from 95.9 to 96.4%). RLE has a slightly higher sensitivity (96.4%) than the TMM and UQ methods while having a tradeoff of lower specificity. When using the recently proposed QL F-test, the results show that UQ-pgQ2 has the highest specificity (58.7% compared to the others arranged from 24.5 to 28.0%) and the highest sensitivity (99.7% compared to the others 99.2%). TMM with a QL F-test has a slightly higher specificity (28%) than RLE/UQ (24.5%). Although a t-test for DEGs analysis in RNA-seq studies is not commonly used due to the distribution of the read counts in RNA-seq data following a negative binomial [26, 49], the voom-limma package has been recently proposed [29] and was reported to have good control of FDR, but low power for small sample size [36, 37]. Therefore, it is interesting to examine the results from a t-test using log-transformation of read counts following one of the four normalization methods. As expected, the results show there is no difference between the UQ and UQ-pgQ2 methods since the median scaling factor estimated for each gene across samples in UQ-pgQ2 was canceled while applying a t-test [50]. Although UQ/UQ-pgQ2 performed relatively better than TMM and RLE, with a specificity of 48.7%, there was a tradeoff with lower power of 93.1%, consistent with previous reports [36, 37]. The results also suggested a t-test is not a better choice for the TMM and RLE methods compared to other tests such as a Wald test or an exact test/QL F-test.
Overall, for this comparison study of the four test statistics (the exact test/QL F-test, Wald test and t-test), the results from MAQC2 data demonstrated that UQ-pgQ2 and TMM combined with an exact test/Wald test performed much better than using a QL F-test and t-test in terms of sensitivity/power and specificity/FDR while UQ and RLE were varied.
The results from an additional analysis of MAQC3 with five replicates had similar conclusions for UQ-pgQ2 normalization (Table 2). Briefly, UQ-pgQ2 with an exact test was the best choice and achieved the highest specificity among the four normalization methods for all four test statistics. The results also show that all normalization methods combined with a t-test/voom-limma achieved better specificity than a Wald test and QL F-test while all the methods have a sensitivity close to or above 90% with a tradeoff of lower power than others. Thus, the results using MAQC3 data suggested that an exact test for UQ-pgQ2 or a t-test from voom-limma seems to control the FDR better than other methods when sample sizes or replicates are relatively large.
Finally, the results from the analysis of both MAQC datasets suggested the four normalization methods combined with the three test statistics (exact test, QL F-test and Wald test) can achieve a great sensitivity/power while a t-test from voom-limma has relatively lower power with unstable performance on the control of FDR. Although the UQ-pgQ2 method performed relatively better for controlling FPs, all normalization methods have a problem maintaining the actual FDR below the nominal level of 0.05, which agrees with previous reports [18, 23, 33].
Within-group analysis of real cancer datasets for detecting FPs given a desired sample size
A type I error rate and FDR are the most important performance measures for evaluating DEG analysis methods. The large number of replicates from TCGA human cancer datasets including non-small cell lung cancer with adenocarcinoma subtype (AdLC), ovarian cancer (OC) and triple negative breast cancer (BC) allows us to perform within-group analysis of FPs for estimation of a type I error rate. The four normalization methods (TMM, RLE and UQ and UQ-pgQ2) combined with the exact test, QL F-test or Wald test were compared given a desired sample size of replicates in a single group. The two synthesized groups with an equal and desired sample size were randomly subsampled from the same cancer subtype. Under the null hypothesis, the genes between the two synthesized groups in this study are not expected to be differentially expressed. Thus, the DE genes identified are defined as FPs. Given a FDR cutoff of 0.05 and an absolute value of FC cutoff at 2 as a conventional way for identifying DEGs, the FPR (a fraction of DEGs) and the number of FPs identified are illustrated in Fig. 1 and Additional file 1: Figure S1, respectively. Although the FPR for all the four normalization methods based on the three datasets are below 0.05, the performance of these methods are significantly different.
False positive rates estimated via intra-group analysis of AdLC, OC and TNBC data. Illustrated are the fractions of FPs estimated from the RLE (pink), TMM (green), UQ (blue) and UQ-pgQ2 (purple) normalization using the exact test, QL F-test or Wald test for sample sizes of 5, 10, 15, 20, 25, 30, 35 and 40. The plots are based on AdLC data (a-c), OC data (d-f) and TNBC data (g-i)
First, using an exact test/QL F-test, we found that the FPR in Fig. 1 increases as the sample size for all the methods increases from 5 to 40 in the three cancer datasets (Fig. 1a, b, d,e, g and h). However, not unexpectedly, this pattern was not observed when a Wald test was used. With a Wald test, higher FPRs are observed when sample sizes are five and they tend to decrease at larger sample sizes of 10, 15, 20, 25, 35 and 40 (Fig. 1c, f and i), but the FPR for different sample sizes varies (Fig. 1c).
Second, we found that the exact test at a sample size of five can achieve a smaller value of FPRs than a Wald test for all the methods (RLE in pink, TMM in green, UQ in blue and UQ-pgQ2 in purple). This suggests that when a sample size is small, an exact test is more conservative than a Wald test. Moreover, the QL F-test combined with any of four normalization methods at sample sizes of 5, 10 and 15 can achieve the smallest FPR compared to the other two tests (Fig. 1b, e and h). However, when a sample size becomes large (n > 15), a Wald test for RLE, TMM and UQ is more conservative than choosing the exact test or QL F-test.
Third, in this study, the differences among RLE, TMM and UQ normalization methods are relatively small and varied. We found that the two-step normalization method UQ-pgQ2, consistently performed better than the others by achieving the smallest FPR and number of FPs given a desired sample size in all scenarios (Fig. 1 and Additional file 1: Figure S1). Overall, the results illustrate that a QL F-test with a UQ-pgQ2 may be the best option for DEG analysis when FDR is more important to be considered. These observations are consistent for the three real datasets.
The effect of sequencing read depth of OC samples on the analysis of FPs from the normalization methods and test statistics
Next, we examined whether the read depth in a RNA-seq study affects the number of FPs for the normalization and test statistical methods given a desired sample size. The read depths of the 379 OC samples range from 19 to 157 million reads. Two new datasets were generated by simply reducing the read depth of the OC samples to one-third or half. Thus, we obtained one dataset with the read depth in the range of 12.8 to 104 million reads and the other with a range of 9.6 to 78.6 million reads. Given an FDR cutoff of 0.05 and an absolute FC cutoff of 2, the FPR estimated from the number of FPs identified in Additional file 1: Figure S2 and S3 is illustrated in Figs. 2 and 3.
False positive rates estimated via intra-group analysis of 379 OC samples with different read depths. Illustrated are the fractions of FPs estimated from the RLE, TMM, UQ and UQ-pgQ2 normalization with the exact test, QL F-test or Wald test for sample sizes of 5, 10, 15, 20, 25, 30, 35 and 40. The plots are based on a read depth from 19 to 157 million (a-c), from 12.8 to 104 million (d-f), and from 9.6 to 78.6 million (g-i)
False positive rates estimated from three test statistics via intra-group analysis of OC samples. Illustrated are the fractions of FPs estimated from the exact test/QL F-test and Wald test combined with the RLE, TMM and UQ-pgQ2 normalization methods, based on the read depths from 19 to 157 million (a-c), from 12.8 to 104 million (d-f) and from 9.6 to 78.6 million (g-i)
Within-group analysis of three sets of data with different read depths revealed that the FPR from higher read depths (19-157 M and 12.8-104 M) in Fig. 2a to 2f is slightly larger than those with smaller read depths (9.6–78.6 M) in Fig. 2g to 2i regardless of normalization methods and sample sizes. However, the difference for the samples with the read depths between 19 and 157 M and 12.8-104 M (a smaller change) is varied and very small. Regardless of read depth, similar patterns are observed between Figs. 1 and 2. Overall, UQ-pgQ2 method is more conservative than the others in most of scenarios given a desired sample size and statistical test. However, Fig. 2 shows that UQ-pgQ2 combined with a Wald test in read depth of 9.6 to 78.6 million is more liberal at the sample size of five.
Figure 3 and Additional file 1: Figure S3 further demonstrate the difference between three test statistics (the exact test in pink, QL F-test in green and Wald test in blue) and three normalization methods. The FPR increases as sample sizes increase while using the exact test and QL F-test, but the impact of FPR by the sequencing read depth are very small. For a Wald test, the FRP is larger than the one from other tests when the sample size is five. Moreover, the FPR from RLE and TMM combined with the three tests are similar (Fig. 3a, b, d, e, g and h). In contrast, UQ-pgQ2 from the three tests (Fig. 3c, f and i) can achieve lower FPR compared to other normalization methods.
The effect of sequencing read depths from simulated data on the analysis of FPs given a desired sample size
Each of the six simulated datasets contains 122 samples with a desired mean read depth of 30, 40 and 50 million reads with a standard deviation (SD) of 3 and 5 million reads, respectively. In this study, we examined whether the simulated data with a desired read depth and SD affects the number of FPs/FPRs from different normalization and test statistical methods given a desired sample size. Given a FDR cutoff of 0.05 and an absolute value of FC cutoff at 2, the results are illustrated in Fig. 4 and Additional file 1: Figure S4.
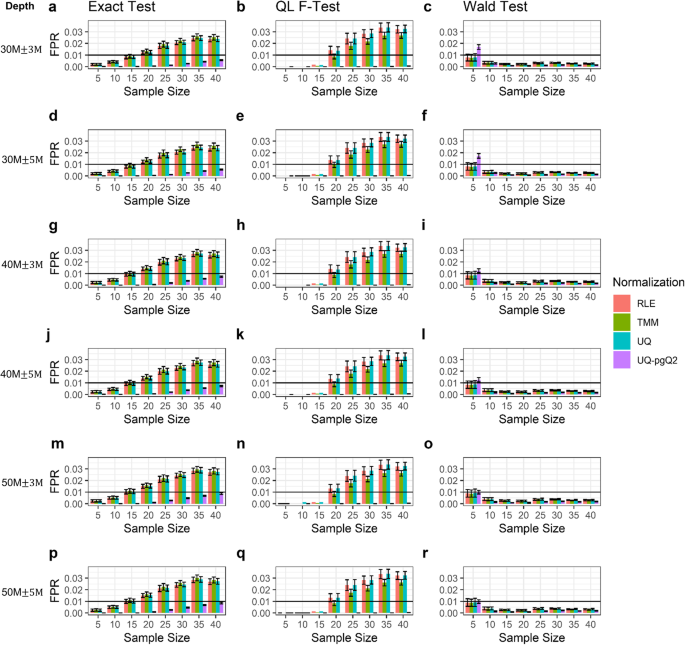
False positive rates estimated from simulated data with variable read depths given fixed sample sizes. Illustrated are the fractions of FPs estimated from the RLE, TMM, UQ and UQ-pgQ2 normalization methods using the exact test, QL F-test or Wald test for sample sizes of 5, 10, 15, 20, 25, 30, 35 and 40. Plots are based on 122 simulated data with the mean read depth of 30 million and SD of 3 (a-c) or 5 million (d-f); a mean read depth of 40 million and SD of 3 (g-i) or 5 million (j-l); a mean read depth of 50 million reads and SD of 3 (m-o) or 5 million (p-r)
Overall, the results from the simulated data are similar to the ones illustrated in Figs. 1 and 2. When the read depths increase from 30 to 50 million reads (Fig. 4a, d, g, j, m and p), the FPR from an exact test slightly increases, which is consistent with the observation from real data (Fig. 1). But, these patterns were not observed when using the Wald test and QL F-test. We also observed that UQ-pgQ2 (purple) performed the best with the lowest FPR while TMM (green) performed the worst with the largest FPR using an exact test. For a Wald test, UQ-pgQ2 performed the worst for a sample size of five and performed the best for sample sizes of 10 or larger while the other methods performed similarly. For a QL F-test, all the methods perform similarly by achieving very small FPR for a given sample size of 5, 10 and 15. When the sample size is 20 or larger, similar results are observed except TMM combined with the QL F-test can achieve a smaller FPR than RLE and UQ methods. Taken together, with a QL F-test, UQ-pgQ2 outperformed the other methods by achieving the smallest value of FPR regardless of the sample sizes and sequencing read depth.
Between-group analysis for identifying DEGs from cancer samples versus the normal controls given a desired sample size
Finally, we evaluated the four normalization methods with three test statistics based on the number of significant DEGs identified using the paired BC data (117 BC and 112 control) and lung cancer data (535 AdLC and 59 normal control). Given a desired sample size, the cancer and control groups were subsampled from BC/AdLC and their control samples for 50 times, respectively. The number of DEGs detected from each method with a desired sample size is the average of the number of DEGs while bootstrapping for 50 times. Given an FDR cutoff of 0.05 and an absolute FC cutoff of 2, the number of DEGs identified is illustrated in Fig. 5.
DEGs identified from the four normalization methods for a balanced two-groups comparison. Illustrated are the number of DEGs identified in 117 TNBC and 112 normal control samples (a, c, e), and 535 AdLC and 59 normal control samples (b, d, f). Analysis was accomplished using the exact test (a, b), QL F-test (c, d) and Wald test (e, f) listed
First, we found that the number of DEGs increases as the sample size increases from 5 to 30 for each method (Fig. 5a-f). As the sample size increases from 30 to 40, the number of DEGs varies either slightly increasing (Fig. 5a-d) or decreasing (Fig. 5e, f).
Second, given sample sizes of 15 or more, an exact test combined with one of the four normalization methods detected more DEGs than the QL F-test and Wald test in the most scenarios, which indicates that the Wald test and QL F-test are more conservative than the exact test while the exact test has more power than others. However, given a sample size of five, the Wald test can identify more DEGs than the exact test and QL F-test.
Third, comparing the four normalization methods, UQ-pgQ2 (purple) is more conservative than the others in the most scenarios except the QL F-test for AdLC data, where UQ-pgQ2 achieved the highest detection power for given sample sizes of 10 or larger (Fig. 5d). However, given a sample size of five, the number of DEGs detected from UQ-pgQ2 combined with the exact test or QL F-test is small (Fig. 5a-c). This observation suggests that RLE, TMM and UQ combined with the exact test or QL F-test from edgeR are the better choice for achieving a better detection power than UQ-pgQ2 for a small sample size such as five and the number of DEGs identified from UQ-pgQ2 looks more reasonable given a sample size ≥10. Since we do not know the number of true positives and true negatives, we assume the method detecting the highest number of DEGs may have the highest sensitivity or detection power.
Discussion
Some previous studies comparing normalization methods have reported that both DESeq2 and edgeR with an exact test/likelihood ratio test failed to maintain the actual FDR below the nominal level of 0.05, suffering from being” oversensitive” in some cases [14, 18, 23, 37, 51]. Our recently proposed normalization method, UQ-pgQ2, combined with an exact test had a better performance than the others in terms of controlling the type I error rate and FDR. However, in that previous study, sample sizes (replicates) for a two-group comparison were smaller than six in many cases. Currently, common sample sizes in RNA-seq studies can range from a minimum of 3 up to several hundreds of biological replicates. It is known that a Wald test can be used for testing a hypothesis of a parameter when sample sizes are usually larger than 30 based on an asymptotic theorem. Recently, edgeR provided a QL F-test which was recommended for studies with a small number of replicates in RNA-seq data. A study of single-cell differential expression analysis observed that edgeR/QLF performed well after filtering out lowly expressed genes [52]. Thus, a comprehensive comparison of QL F-test and other test statistics combined with newly developed normalization methods (UQ-pgQ2 or voom-limma) for DEG analysis of bulk RNA-seq data deem to be needed. Furthermore, one outstanding question is whether the Wald test, QL F-test or t-test combined with a normalization method performs better than an exact test used in the previous study in terms of controlling FDR. Another question is what is the best combination between normalization and test statistical methods for the control of type I error rate given a desired sample size. To address these issues, we focused on four normalization methods and three test statistics using both real RNA-seq datasets and simulated data given sample sizes at 5, 10, 15, 20, 25, 30, 35 and 40.
Initially, we used two benchmark MAQC datasets for the DEGs analysis. The results from MAQC datasets show that UQ-pgQ2 combined with an exact test can achieve the highest specificity with the sensitivity higher than 90%. We also observed that edgeR with TMM normalization performed better than RLE and UQ with the Wald test or QL F-test in terms of sensitivity and specificity using the small samples sizes of MAQC datasets.
Next, we used an intra-group approach to calculate the number of FPs and FPR from null datasets generated from real data and simulated data. We compared these methods by taking into consideration of sample sizes and read depths. The results from this approach have shown that in general the QL F-test combined with any one of four normalization methods (RLE, TMM, UQ and UQ-pgQ2) performs the best for achieving the lowest value of FPRs when the sample size is small (≤15). However, when the sample size is large, UQ-pgQ2 combined with the QL F-test performs the best and the Wald test performs much better than an exact test and QL F-test with FPR below 0.01. Moreover, we found the read depths from simulated data with 30 to 50 million reads have a minimal impact on detection power and FDR. Furthermore, comparing DEG analysis of BC and AdLC suggests that the RLE, TMM and UQ combined with an exact test or a Wald test have higher sensitivity or power than the UQ-pgQ2 method. However, these methods may suffer oversensitivity when the sample sizes are large. In addition, the results from AdLC show UQ-pgQ2 combined the QL F-test achieves higher detection power than other combined methods, but this observation is not consistent with the one from BC datasets.
Furthermore, it is important to note that the evaluated methods may not be applicable to all type RNA-seq data. For example, single-cell RNA sequencing (scRNA-seq) data has been increasingly used to assess different cell states and cell types such as stem cell, neuron cell and cancer cells [53,54,55]. A DEG analysis between different cells in scRNA-seq can help to uncover driver genes in cancer research [56]. Due to technical limitations, scRNA-seq data generally have low library sizes resulting in a large fraction of ‘dropout’ events as well as huge heterogeneity, which introduces a major challenge in identification of DEGs. Given the special characteristics, many new methods have been developed especially for DE analysis of scRNA-seq data [57,58,59,60]. A few studies have compared DEG analysis tools for scRNA-seq data and found that existing methods for analysis of bulk RNA-seq data perform as well as, or not worse than, those specifically developed for scRNA-seq data in terms of the power and FDR [52, 61]. A recent study has conducted DEG analysis for a comparison of eleven tools and two of them (edgeR and DESeq2) were designed originally for bulk RNA-seq [62]. In this study, a high disagreement among these tools in calling DEGs were identified due to a trade-off between the sensitivity and FDR. It also reported that current methods developed for scRNA-seq did not show better performance compared to edgeR and DESeq2, which is consistent with the findings from previous studies. Thus, it is reasonable for us to mention here whether the newly developed normalization methods such as UQ-pgQ2 and Med-pgQ2 can be used for DEG analysis in scRNA-seq data for a control of FDR.
Finally, our study has some limitations. First, this work is limited to gene-level analysis and balanced designs. Second, the per gene normalization method, UQ-pgQ2, is only used for the DEG analysis among the groups and is not applicable for comparing genes within a group. Third, we used within group analysis of several cancer subtypes to identify DEGs and we assumed the DEGs as false positives that were used to calculate FPRs. Although this approach has been used to estimate the type I error or FPs for the comparison of normalization and test statistical methods in several studies [32, 37, 46], these identified genes may contain some TPs due to the variation between cancer patients. However, this limitation can be offset by the two benchmark MAQC RNA-seq data. Finally, since voom-limma with a t-test used for DEGs analysis in bulk RNA-seq data, we need to address here that per gene normalization in UQ-pgQ2 would not alter the DEG results due to the invariant property of t-test for the linear transformations of gene counts across samples.
Conclusions
Taken together, we found the UQ-pgQ2 method with an exact test is the best choice for DEG analysis in terms of controlling false positives when using the benchmark MAQC datasets. However, based on an intra-group analysis of real data and simulated data, we found UQ-pgQ2 combined with a QL F-test outperforms other methods by achieving the smallest value of FPR, and a Wald test from DESeq2 can achieve a FPR below 0.05 when sample sizes are large. We observed that the RLE, TMM and UQ normalization methods combined with the Wald or exact test/QL F-test performed similarly and read depths have minimal impact on detection of DEGs from the analysis of simulated data. We hope this new finding can serve a guide for researchers to properly choose the normalization and test statistical methods for identifying DEGs while taking sample sizes into the consideration. As scRNA-seq technology emerges, UQ-pgQ2 combined with the QL F-test may be useful for DEG analysis of scRNA-seq data for controlling FDR, but an evaluation should be conducted in the future.
Methods
Data sources
Microarray quality control project (MAQC) RNA-seq data
Two benchmark RNA-seq datasets (MAQC2 and MAQC3) were used for evaluation [18, 34, 46]. These datasets have two conditions: human brain reference RNA (hbr) and universal human reference RNA (uhr). MAQC2 RNA-seq data contains two replicates in each condition (hbr1, hbr2, uhr1 and uhr2). The hbr1 / uhr1 and hbr2 / uhr2 samples were prepared and sequenced in different labs. MAQC3 RNA-seq data contains five replicates in each condition.
TaqMan qRT-PCR data
In the MAQC project, a benchmark PCR dataset of 1044 genes was used for validation. Detailed information for processing and analyzing this data has been previously described [17, 18, 38, 47]. Briefly, we identified 388 genes as true positives and 143 genes as true negatives to evaluate these methods.
Human cancer RNA-seq datasets from TCGA
Three types of human BC, OC and LC data were downloaded from TCGA website [48]. In this study, we used one subtype from each cancer type. The 122 TNBC, 379 OC, 523 AdLC and 59 normal control samples were extracted using an R script (v3.5.2). For DEG analysis, 117 BC and 112 paired normal control samples were extracted. Lowly expressed genes in each TCGA dataset were filtered out when the zero counts across samples were greater than 50% across the samples. The remaining genes were used for the downstream analysis. These datasets are available in Additional file 2.
Simulated data using Monte Carlo method
The simulated data is based on the 122 TNBC samples with a range of total read depth between 23.8 and 98.7 million reads. For our simulation, let G be the total number of genes (G = 30,831) with desired sample sizes (n = 5, 10, 15, 20, 25, 30, 35 and 40) denoted as the number of replicates. Let Rig be the read counts in sample i and gene g and Ni be the total number of reads (read depth) in sample i as estimated from the 122 TNBC samples. Let pig be the proportion for the gene g in sample i, where \( {\hat{p}}_{ig} \) was estimated from Rig divided by Ni. For simulating read depths, we modeled the independent and desired read depth Ni (i = 1, …, ni) to follow a normal distribution with a mean of 30, 40 and 50 million reads, and standard deviation of 3 million reads for scenario one and 5 million reads for scenario two for each N,. The resulting distributions are Ni ∼ N (30, 32) and Ni ∼ N(30, 52); Ni ∼ N(40, 32) and Ni ∼ N(40, 52); and Ni ∼ N(50, 32) and Ni ∼ N(50, 52), respectively.
As an example, in the case of generating samples with a mean read depth of 30 million reads and a standard deviation of 3 million reads given a desired sample size of five, first, ten samples for the two groups were randomly selected from the set of 122 TNBC cases. We estimated \( {\hat{p}}_{ig} \) for each sample and generated read depths based on Ni ∼ N(30, 32) using the Monte Carlo method. Finally, the raw counts for each sample were generated using multinomial distribution given a desired Ni and a vector of \( {\hat{p}}_{ig} \) for sample i. This procedure was repeated 50 times given desired sample size of 5, 10, 15, 20, 25, 30, 35 or 40 combined with Ni ∼ N(30, 32). Thus, the data in the two groups was generated in the scenario of Ni ∼ N(30, 32).
Sequence alignment and extraction of gene counts
The raw Sequence Read Archive (SRA) files for the MAQC2 and MAQC3 with two conditions were converted to .fastq files and then aligned to the human hg38 reference genome using STAR (v2.6.0c) [11] and the Ensembl hg38 annotation gtf file (GRCh38.82). The mapped counts for 60,483 genes per sample were extracted using HTSeq-scripts-count (version 2.7.5). Lowly expressed genes with zero counts across all the samples were further filtered. The MAQC datasets are available in Additional file 2.
Normalization methods and software packages
The normalization methods, statistical tests and software packages for the DEG analysis between the two-group comparisons are summarized in Table 3. The R code used in our analysis is available in Additional file 3.
Normalization methods
Four methods (RLE, TMM, UQ and UQ-pgQ2) were used to normalize RNA-seq data. In this study, RLE and TMM normalizations were implemented in DESeq2 and edgeR, respectively [25, 63]. UQ and UQ-pgQ2 were implemented using an R script [18, 23, 28]. UQ-pgQ2 uses a two-step approach for normalization [18]. Briefly, assuming G genes and m samples, the scaling factor for UQ normalization is calculated from the 75th percentile (Q3) of the counts for each sample after removing genes with zero counts. Gene g in sample j is scaled by the UQ scaling factor and then multiplied by the mean of the scaling factors from m samples. Therefore, \( {X}_{gj}^{UQ} \) is gene g in sample j with UQ normalized counts. Then, UQ-normalized gene g is further scaled by its median (Q2) of read counts across m samples and then multiplied by 100. Therefore, \( {X}_{gj}^{UQ- pgQ2} \) is gene g in sample j and normalized by UQ-pgQ2 method.
Software packages and test statistics used for DEGs analysis
The exact test, QL F-test and Wald test were used for the detection of DEGs. In this study, we used the exact test and QL F-test implemented in edgeR (v3.24.3) [45]. A Wald test implemented in DESeq2 (v1.22.2) was reported to improve the sensitivity compared with an exact test implemented in DESeq [25].
Log-transformed Wald test in a NB distribution
The Wald statistical test is an asymptotic test based on the normal approximation, which utilizes the large-sample properties of maximum likelihood estimation (MLE). In DESeq2, the read counts Kij in gene i sample j is modelled by a generalized linear model (GLM) of the NB family with a log link:
Kij~NB(mean = uij, Dispersion = αi), uij = sijuij and log \( {q}_{ij}=\sum r{x}_{jr}{\beta}_{ir} \). The sij = sj is the size factor used to normalize the gene read counts in sample j and the qij is the true expression of gene i [25].
For a GLM with two conditions and a single gene in sample j, the log-transformed Wald test has been described in several studies [64,65,66]. Briefly, the treatment and control group indicators (xr) take the value 1 and 0, respectively, resulting \( \frac{q_1}{q_0}={e}^{\beta_1} \) . For the differential expression gene analysis, the ratio ρ =q1/q0 (a fold change) is used for a hypothesis test. Testing the hypothesis of qr: H0: ρ = 1 vs. H1: ρ ≠ 1 is equivalent to H0: β1 = 0 vs. H1: β1 ≠ 0 in GLM. The two-sided Wald test is defined as \( \mid {Z}_w\mid =\frac{\left|{\hat{\beta}}_1\right|}{\sqrt{Var\left({\hat{\beta}}_1\right)}} \), where the \( Var\left({\hat{\beta}}_1\right) \) is estimated from the variance-covariance matrix of (β0 ,β1), which is the inverse of the Fisher information matrix of \( {I}_{n_0{n}_1}\left({\beta}_0\ \mathrm{and}\ {\beta}_1\right) \) asymptotically. To reject the null hypothesis, \( \left|{Z}_w\right|>{Z}_{1-\frac{\alpha }{2}} \) is defined. Thus, this gene with p-value less than 0.05 is called a DEG. For testing multiple genes simultaneously, the p-values are further corrected using Benjamini-Houchberg FDR method [67].
Exact test in a NB distribution
Since RNA-seq data are read counts, an exact test has been implemented similarly in DESeq and edgeR [26, 68]. For a gene in a two-group comparison, the exact test has been described by several studies [18, 26, 69] . Briefly, Yij is denoted the normalized read counts of the single gene in condition i = A and B, and replicate j = 1, …, ni. Then, the distributions of \( {Y}_{ij}\ \mathrm{and}\ \sum \limits_j^{n_i}{Y}_{ij} \) are assumed to follow a negative binomial distribution with an expected mean ui and dispersion ϕ expressed as:
Ygij~Ygij~NB(ugi, ϕ), \( \sum \limits_j^{n_i}{Y}_{ij}\sim NB\left({n}_i\cdotp {u}_i,\frac{\upphi\ }{n_i}\right),E\left(\sum \limits_j^{n_i}{Y}_{ij}\right)={n}_i\cdotp {u}_i \), and Var \( \left(\sum \limits_j^{n_i}{Y}_{ij}\right)={n}_i\cdotp {u}_i+{n}_i\cdotp {u}_i^2\cdotp \upphi \). The null hypothesis H0 : uA = uB is to identify DEGs between conditions A and B and the total normalized counts in each condition are \( {Y}_A=\sum \limits_j^{n_A}{Y}_{Aj} \) and \( {Y}_B=\sum \limits_j^{n_B}{Y}_{Bj}. \) The total counts of two conditions for the gene are YS = YA + YB.
Since YA and YB are assumed to be independent, the joint probability of P(YA = yA, YB = yB ) under H0 is P(YgA = ygA) × P(YgB = ygB). Thus, the p-value from an exact test [26] is calculated by summation of the probability of a pair of P(a, b) that is less than or equal to the observed P(yA, yB) given that the overall summation of P(a, b). The pair of variables a and b are defined as a = 0, …, YS and b = YS − a. Then the p-value for gene g is
p. value = P1/P2, where
and
The p-value is further adjusted for multiple test correction using the Benjamini-Hochberg FDR methodology. A study reported that the exact test performed better for achieving a smaller FDR than a Wald and likelihood ratio tests when the sample size is small [49].
Intra-group analysis to identify the number of FPs given a desired sample size
To compare the methods, we utilized the desired number of replicates (5, 10, 15, 20, 25, 30, 35 and 40) to estimate the number of false positives and the corresponding FPR, a fraction of DEGs via intra-group analysis. Given sample sizes, two groups are generated by randomly subsampling from a single cancer group. Since the samples originate from the same condition, the number of DEGs expected in such two-group comparisons should be zero. Thus, by this assumption, any DEGs would be defined as false positives that were further used for estimating FPR under a null hypothesis [32, 37, 52, 70].
Bootstrap differential expression analysis of cancer versus normal control samples given a desired sample size
Given desired sample sizes of 5, 10, 15, 20, 25, 30, 35 and 40, the two-group data (cancer and normal control) were randomly subsampling from BC/AdLC and the normal samples, respectively. Subsequently, the cancer and control groups were normalized by one of the four normalization methods used in this study. The DEG analysis of the normalized data was performed with the aid of edgeR, DESeq2 and limma tools. The results from these packages include the log2 transformed fold change, p-values and Benjamini-Hochberg FDRs. In this study, we defined significant DEGs using an absolute value of log2 FC cutoff at one (|FC| = 2) and a FDR cutoff at 0.05. The bootstrapping process of running each DEG algorithm was iteratively repeated 50 times. The mean number of DEGs corresponding to the standard error was imputed from the 50 iterations.
Abbreviations
- AdLC:
-
Human non-small cell lung cancer with adenocarcinoma subtype
- BC:
-
Human breast cancer
- DE:
-
Differential expression
- DEGs:
-
Differentially expressed genes
- FC:
-
Fold change
- FDR:
-
False discovery rate
- FP:
-
False positive
- FPKM:
-
Fragments Per Kilobase per Million mapped fragments
- FPR:
-
False positive rate
- hbr:
-
Human brain reference RNA
- LC:
-
Human lung cancer
- LRT:
-
Likelihood ratio test
- MAQC:
-
Microarray quality control project
- Med-pgQ2:
-
per-gene Q2 normalization following per-sample median global scaling
- MLE:
-
Maximum likelihood estimation
- NB:
-
Negative binomial distribution
- OC:
-
Human ovarian cancer
- PPV:
-
Positive predict value
- Q:
-
Full Quantile
- Q2:
-
The 50th percentile
- Q3:
-
The 75th percentile
- QL F-test:
-
Quasi-likelihood F test
- qRT-PCR:
-
Quantitative Real-Time polymerase chain reaction
- RC:
-
Raw counts
- RLE:
-
Relative log estimate
- RNA-seq:
-
High-throughput RNA sequencing
- ROC:
-
Receiver Operating Characteristic
- RPKM:
-
Reads Per Kilobase per Million mapped reads
- RSEM:
-
RNA-seq by expectation-maximization
- RUV:
-
Remove unwanted technical variation
- scRNA-seq:
-
Single cell RNA sequencing
- SD:
-
Standard deviation
- SRA:
-
Sequenced read archive
- TC:
-
Total counts
- TCGA:
-
The Cancer Genome Atlas
- TMM:
-
Trimmed-mean M values
- TNBC:
-
Triple negative breast cancer
- TP:
-
True positive
- uhr:
-
Universal human reference RNA
- UQ:
-
Upper Quartile (Q3)
- UQ-pgQ2:
-
per-gene Q2 normalization following per-sample upper-quartile global scaling at 75 percentile
References
Wang Z, Gerstein M, Snyder M. RNA-Seq: a revolutionary tool for transcriptomics. Nat Rev Genet. 2009;10(1):57–63.
Marguerat S, Bahler J. RNA-seq: from technology to biology. Cell Mol Life Sci. 2010;67(4):569–79.
Robertson G, Schein J, Chiu R, Corbett R, Field M, Jackman SD, et al. De novo assembly and analysis of RNA-seq data. Nat Methods. 2010;7(11):909–12.
Schliebner I, Becher R, Hempel M, Deising HB, Horbach R. New gene models and alternative splicing in the maize pathogen Colletotrichum graminicola revealed by RNA-Seq analysis. BMC Genomics. 2014;15(1):842.
Canovas A, Rincon G, Islas-Trejo A, Wickramasinghe S, Medrano JF. SNP discovery in the bovine milk transcriptome using RNA-Seq technology. Mamm Genome. 2010;21(11–12):592–8.
Piskol R, Ramaswami G, Li JB. Reliable identification of genomic variants from RNA-seq data. Am J Hum Genet. 2013;93(4):641–51.
Wang L, Feng Z, Wang X, Wang X, Zhang X. DEGseq: an R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics. 2010;26(1):136–8.
Ellard S, Patrinos GP, Oetting WS. Clinical applications of next-generation sequencing: the 2013 human genome variation society scientific meeting. Hum Mutat. 2013;34(11):1583–7.
Byron SA, Van Keuren-Jensen KR, Engelthaler DM, Carpten JD, Craig DW. Translating RNA sequencing into clinical diagnostics: opportunities and challenges. Nat Rev Genet. 2016;17(5):257–71.
Langmead B, Salzberg SL. Fast gapped-read alignment with bowtie 2. Nat Methods. 2012;9(4):357–9.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29(1):15–21.
Trapnell C, Pachter L, Salzberg SL. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics. 2009;25(9):1105–11.
Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015;12(4):357–60.
Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013;14(4):R36.
Zeng W, Mortazavi A. Technical considerations for functional sequencing assays. Nat Immunol. 2012;13(9):802–7.
Robinson MD, Oshlack A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 2010;11(3):R25.
Bullard JH, Purdom E, Hansen KD, Dudoit S. Evaluation of statistical methods for normalization and differential expression in mRNA-Seq experiments. BMC Bioinformatics. 2010;11:94.
Li X, Brock GN, Rouchka EC, Cooper NGF, Wu D, O'Toole TE, et al. A comparison of per sample global scaling and per gene normalization methods for differential expression analysis of RNA-seq data. PLoS One. 2017;12(5):e0176185.
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods. 2008;5(7):621–8.
Marioni JC, Mason CE, Mane SM, Stephens M, Gilad Y. RNA-seq: an assessment of technical reproducibility and comparison with gene expression arrays. Genome Res. 2008;18(9):1509–17.
Trapnell C, Williams BA, Pertea G, Mortazavi A, Kwan G, van Baren MJ, et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol. 2010;28(5):511–5.
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and cufflinks. Nat Protoc. 2012;7(3):562–78.
Dillies MA, Rau A, Aubert J, Hennequet-Antier C, Jeanmougin M, Servant N, et al. A comprehensive evaluation of normalization methods for Illumina high-throughput RNA sequencing data analysis. Brief Bioinform. 2013;14(6):671–83.
Smyth GK. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol. 2004;3:Article 3.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15(12):550.
Anders S, Huber W. Differential expression analysis for sequence count data. Genome Biol. 2010;11(10):R106.
Risso D, Ngai J, Speed TP, Dudoit S. Normalization of RNA-seq data using factor analysis of control genes or samples. Nat Biotechnol. 2014;32(9):896–902.
Risso D, Schwartz K, Sherlock G, Dudoit S. GC-content normalization for RNA-Seq data. BMC Bioinformatics. 2011;12:480.
Law CW, Chen Y, Shi W, Smyth GK. voom: precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 2014;15(2):R29.
Li B, Dewey CN. RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinformatics. 2011;12:323.
Patro R, Mount SM, Kingsford C. Sailfish enables alignment-free isoform quantification from RNA-seq reads using lightweight algorithms. Nat Biotechnol. 2014;32(5):462–4.
Li X, Rouchka EC, Brock GN, Yan J, O'Toole TE, Tieri DA, et al. A combined approach with gene-wise normalization improves the analysis of RNA-seq data in human breast cancer subtypes. PLoS One. 2018;13(8):e0201813.
Kvam VM, Liu P, Si Y. A comparison of statistical methods for detecting differentially expressed genes from RNA-seq data. Am J Bot. 2012;99(2):248–56.
Rapaport F, Khanin R, Liang Y, Pirun M, Krek A, Zumbo P, et al. Comprehensive evaluation of differential gene expression analysis methods for RNA-seq data. Genome Biol. 2013;14(9):R95.
Li P, Piao Y, Shon HS, Ryu KH. Comparing the normalization methods for the differential analysis of Illumina high-throughput RNA-Seq data. BMC Bioinformatics. 2015;16:347.
Soneson C, Delorenzi M. A comparison of methods for differential expression analysis of RNA-seq data. BMC Bioinformatics. 2013;14:91.
Seyednasrollah F, Laiho A, Elo LL. Comparison of software packages for detecting differential expression in RNA-seq studies. Brief Bioinform. 2015;16(1):59–70.
Zhang ZH, Jhaveri DJ, Marshall VM, Bauer DC, Edson J, Narayanan RK, et al. A comparative study of techniques for differential expression analysis on RNA-Seq data. PLoS One. 2014;9(8):e103207.
Lin Y, Golovnina K, Chen ZX, Lee HN, Negron YL, Sultana H, et al. Comparison of normalization and differential expression analyses using RNA-Seq data from 726 individual Drosophila melanogaster. BMC Genomics. 2016;17:28.
Tang M, Sun J, Shimizu K, Kadota K. Evaluation of methods for differential expression analysis on multi-group RNA-seq count data. BMC Bioinformatics. 2015;16:361.
Germain PL, Vitriolo A, Adamo A, Laise P, Das V, Testa G. RNAontheBENCH: computational and empirical resources for benchmarking RNAseq quantification and differential expression methods. Nucleic Acids Res. 2016;44(11):5054–67.
Maza E. In Papyro comparison of TMM (edgeR), RLE (DESeq2), and MRN normalization methods for a simple two-conditions-without-replicates RNA-Seq experimental design. Front Genet. 2016;7:164.
Costa-Silva J, Domingues D, Lopes FM. RNA-Seq differential expression analysis: an extended review and a software tool. PLoS One. 2017;12(12):e0190152.
Spies D, Renz PF, Beyer TA, Ciaudo C. Comparative analysis of differential gene expression tools for RNA sequencing time course data. Brief Bioinform. 2019;20(1):288–98.
Lun AT, Chen Y, Smyth GK. It's DE-licious: a recipe for differential expression analyses of RNA-seq experiments using quasi-likelihood methods in edgeR. Methods Mol Biol. 2016;1418:391–416.
Consortium M, Shi L, Reid LH, Jones WD, Shippy R, Warrington JA, et al. The MicroArray quality control (MAQC) project shows inter- and intraplatform reproducibility of gene expression measurements. Nat Biotechnol. 2006;24(9):1151–61.
Wan L, Sun F. CEDER: accurate detection of differentially expressed genes by combining significance of exons using RNA-Seq. IEEE/ACM Trans Comput Biol Bioinform. 2012;9(5):1281–92.
Grossman RL, Heath AP, Ferretti V, Varmus HE, Lowy DR, Kibbe WA, et al. Toward a shared vision for Cancer genomic data. N Engl J Med. 2016;375(12):1109–12.
Robinson MD, Smyth GK. Small-sample estimation of negative binomial dispersion, with applications to SAGE data. Biostatistics. 2008;9(2):321–32.
Li X, Gu G, Soliman F, Sanders AJ, Wang X, Liu C. The evaluation of durative transfusion of Endostar combined with chemotherapy in patients with advanced non-small cell lung cancer. Chemotherapy. 2018;63(4):214–9.
Guo Y, Li CI, Ye F, Shyr Y. Evaluation of read count based RNAseq analysis methods. BMC Genomics. 2013;14(Suppl 8):S2.
Soneson C, Robinson MD. Bias, robustness and scalability in single-cell differential expression analysis. Nat Methods. 2018;15(4):255–61.
Klein AM, Mazutis L, Akartuna I, Tallapragada N, Veres A, Li V, et al. Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell. 2015;161(5):1187–201.
Usoskin D, Furlan A, Islam S, Abdo H, Lonnerberg P, Lou D, et al. Unbiased classification of sensory neuron types by large-scale single-cell RNA sequencing. Nat Neurosci. 2015;18(1):145–53.
Zheng C, Zheng L, Yoo JK, Guo H, Zhang Y, Guo X, et al. Landscape of infiltrating T cells in liver Cancer revealed by single-cell sequencing. Cell. 2017;169(7):1342–56 e1316.
Myers JS, von Lersner AK, Robbins CJ, Sang QX. Differentially expressed genes and signature pathways of human prostate Cancer. PLoS One. 2015;10(12):e0145322.
Kharchenko PV, Silberstein L, Scadden DT. Bayesian approach to single-cell differential expression analysis. Nat Methods. 2014;11(7):740–2.
Finak G, McDavid A, Yajima M, Deng J, Gersuk V, Shalek AK, et al. MAST: a flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA sequencing data. Genome Biol. 2015;16:278.
Nabavi S, Schmolze D, Maitituoheti M, Malladi S, Beck AH. EMDomics: a robust and powerful method for the identification of genes differentially expressed between heterogeneous classes. Bioinformatics. 2016;32(4):533–41.
Miao Z, Deng K, Wang X, Zhang X. DEsingle for detecting three types of differential expression in single-cell RNA-seq data. Bioinformatics. 2018;34(18):3223–4.
Jaakkola MK, Seyednasrollah F, Mehmood A, Elo LL. Comparison of methods to detect differentially expressed genes between single-cell populations. Brief Bioinform. 2017;18(5):735–43.
Wang T, Li B, Nelson CE, Nabavi S. Comparative analysis of differential gene expression analysis tools for single-cell RNA sequencing data. BMC Bioinformatics. 2019;20(1):40.
Robinson MD, McCarthy DJ, Smyth GK. edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26(1):139–40.
Keene ON, Jones MR, Lane PW, Anderson J. Analysis of exacerbation rates in asthma and chronic obstructive pulmonary disease: example from the TRISTAN study. Pharm Stat. 2007;6(2):89–97.
Zhu H, Lakkis H. Sample size calculation for comparing two negative binomial rates. Stat Med. 2014;33(3):376–87.
Li X, Wu D, Cooper NGF, Rai SN. Sample size calculations for the differential expression analysis of RNA-seq data using a negative binomial regression model. Stat Appl Genet Mol Biol. 2019;18(1). https://doi.org/10.1515/sagmb-2018-0021.
Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc B. 1995;57:289–300.
Robinson MD, Smyth GK. Moderated statistical tests for assessing differences in tag abundance. Bioinformatics. 2007;23(21):2881–7.
Yu D, Huber W, Vitek O. Shrinkage estimation of dispersion in negative binomial models for RNA-seq experiments with small sample size. Bioinformatics. 2013;29(10):1275–82.
Schurch NJ, Schofield P, Gierlinski M, Cole C, Sherstnev A, Singh V, et al. How many biological replicates are needed in an RNA-seq experiment and which differential expression tool should you use? RNA. 2016;22(6):839–51.
Acknowledgements
The authors gratefully acknowledge the reviewers for the insightful suggestions and advice during the revision.
Funding
This research was supported by the National Institutes of Health, P20GM103436 to Dr. Nigel Cooper. The funding bodies played no role in the design of the study and collection, analysis, and interpretation of data and in writing the manuscript.
Author information
Authors and Affiliations
Contributions
XL conducted the simulation, statistical and computational analysis, and drafted the manuscript. ECR and NGFC advised and participated in discussion. XL, TEO, ECR and NGFC revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Additional file 2.
Contains datasets used for the analysis. This zipped file folder contains MAQC2 and MAQC3 raw read counts, cancer raw data files filtered with zero counts (AdLC, OC and TNBC), and description of these data files named Supplementary Material.docx.
Additional file 3.
R code for the analysis.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated.
About this article
Cite this article
Li, X., Cooper, N.G.F., O’Toole, T.E. et al. Choice of library size normalization and statistical methods for differential gene expression analysis in balanced two-group comparisons for RNA-seq studies. BMC Genomics 21, 75 (2020). https://doi.org/10.1186/s12864-020-6502-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12864-020-6502-7